Projects created before 2024-01-08 are available to browse. Unfortunately templates originating from AlphaFold Database (AFDB) will fail for model building.
Please start a new template search to build models based on AFDB templates.
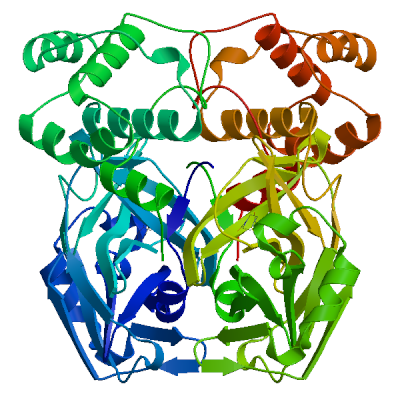
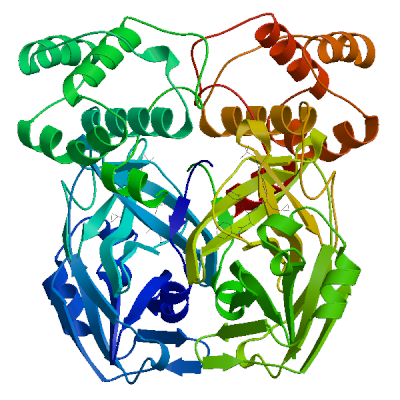
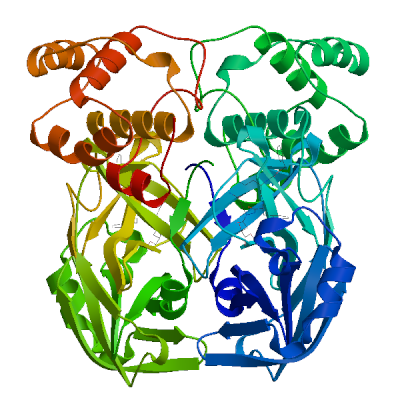
3C-like proteinase nsp5 (3CL-PRO) | P0DTD1 PRO_0000449623
Created: May 5, 2023 at 21:34
Model 02
- Ligands
2 x T8J 2 x 1-[4-(thiophene-2-carbonyl)piperazin-1-yl]ethan-1-oneT8J.4: 11 residues within 4Å:- Chain A: T.25, H.41, C.44, T.45, S.46, M.49, N.142, G.143, S.144, C.145, H.163
4 PLIP interactions:4 interactions with chain A- Hydrophobic interactions: A:T.25
- Hydrogen bonds: A:G.143, A:S.144, A:C.145
T8J.8: 11 residues within 4Å:- Chain B: T.25, H.41, C.44, T.45, S.46, M.49, N.142, G.143, S.144, C.145, H.163
4 PLIP interactions:4 interactions with chain B- Hydrophobic interactions: B:T.25
- Hydrogen bonds: B:G.143, B:S.144, B:C.145
QMEANDisCo Local
QMEAN Z-Scores
Template
5rfv.1.B
3C-like proteinase
PanDDA analysis group deposition -- Crystal Structure of SARS-CoV-2 main protease in complex with PCM-0102306
PanDDA analysis group deposition -- Crystal Structure of SARS-CoV-2 main protease in complex with PCM-0102306
- Seq Identity
- 100.00%
- Coverage
| Biounit Oligo State | Homo-dimer |
| QSQE | 0.95 |
| Method | X-ray, 1.48 Å |
| Seq Similarity | 0.63 |
| Coverage | 1.00 |
| Range | 1-304 |
| Ligand | Added to Model | Description |
|---|---|---|
| ✓ | 1-[4-(thiophene-2-carbonyl)piperazin-1-yl]ethan-1-one | |
| ✓ | 1-[4-(thiophene-2-carbonyl)piperazin-1-yl]ethan-1-one | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE |
Model-Template Alignment
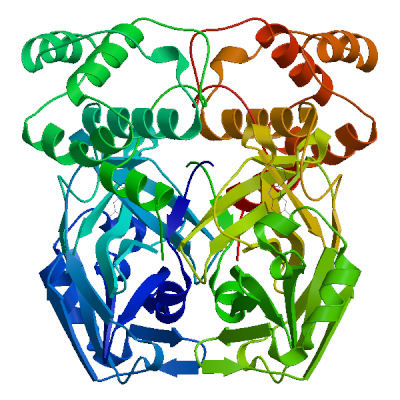
Model 04
- Ligands
2 x HWH 2 x ~{N}-[2-(5-fluoranyl-1~{H}-indol-3-yl)ethyl]ethanamideHWH.4: 11 residues within 4Å:- Chain A: H.41, M.49, H.164, M.165, E.166, L.167, P.168, D.187, R.188, Q.189, T.190
5 PLIP interactions:5 interactions with chain A- Hydrophobic interactions: A:M.165, A:Q.189
- Hydrogen bonds: A:E.166, A:Q.189
- pi-Stacking: A:H.41
HWH.9: 11 residues within 4Å:- Chain B: H.41, M.49, H.164, M.165, E.166, L.167, P.168, D.187, R.188, Q.189, T.190
5 PLIP interactions:5 interactions with chain B- Hydrophobic interactions: B:M.165, B:Q.189
- Hydrogen bonds: B:E.166, B:Q.189
- pi-Stacking: B:H.41
QMEANDisCo Local
QMEAN Z-Scores
Template
5r7z.1.B
3C-like proteinase
PanDDA analysis group deposition -- Crystal Structure of SARS-CoV-2 main protease in complex with Z1220452176
PanDDA analysis group deposition -- Crystal Structure of SARS-CoV-2 main protease in complex with Z1220452176
- Seq Identity
- 100.00%
- Coverage
| Biounit Oligo State | Homo-dimer |
| QSQE | 0.97 |
| Method | X-ray, 1.59 Å |
| Seq Similarity | 0.63 |
| Coverage | 1.00 |
| Range | 1-304 |
| Ligand | Added to Model | Description |
|---|---|---|
| ✓ | ~{N}-[2-(5-fluoranyl-1~{H}-indol-3-yl)ethyl]ethanamide | |
| ✓ | ~{N}-[2-(5-fluoranyl-1~{H}-indol-3-yl)ethyl]ethanamide | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE | |
| ✕ - Not biologically relevant. | DIMETHYL SULFOXIDE |
Model-Template Alignment
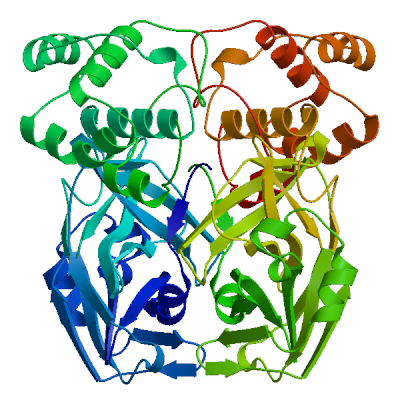
Model 05
QMEANDisCo Local
QMEAN Z-Scores
Template
6lu7.1.B
3C-like proteinase
The crystal structure of COVID-19 main protease in complex with an inhibitor N3
The crystal structure of COVID-19 main protease in complex with an inhibitor N3
- Seq Identity
- 100.00%
- Coverage
| Biounit Oligo State | Homo-dimer |
| QSQE | 0.99 |
| Method | X-ray, 2.16 Å |
| Seq Similarity | 0.63 |
| Coverage | 1.00 |
| Range | 1-306 |
| Ligand | Added to Model | Description |
|---|---|---|
| ✕ - Clashing with protein. | N-[(5-METHYLISOXAZOL-3-YL)CARBONYL]ALANYL-L-VALYL-N~1~-((1R,2Z)-4-(BENZYLOXY)-4-OXO-1-{[(3R)-2-OXOPYRROLIDIN-3-YL]METHYL}BUT-2-ENYL)-L-LEUCINAMIDE | |
| ✕ - Clashing with protein. | N-[(5-METHYLISOXAZOL-3-YL)CARBONYL]ALANYL-L-VALYL-N~1~-((1R,2Z)-4-(BENZYLOXY)-4-OXO-1-{[(3R)-2-OXOPYRROLIDIN-3-YL]METHYL}BUT-2-ENYL)-L-LEUCINAMIDE |
Model-Template Alignment
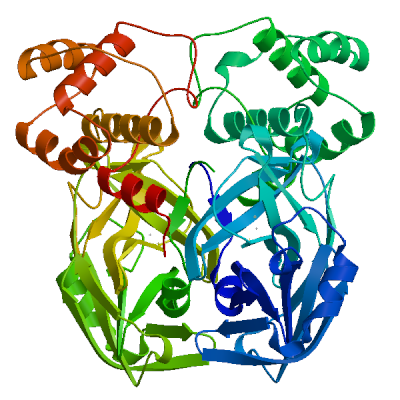
Model 06
- Ligands
2 x DTZ 2 x ZINC(II)HYDROGENSULFIDEDTZ.5: 3 residues within 4Å:- Chain B: H.41, C.145, H.164
2 PLIP interactions:2 interactions with chain B- Metal complexes: B:H.41, B:C.145
DTZ.6: 5 residues within 4Å:- Chain A: T.25, L.27, H.41, C.145, H.164
2 PLIP interactions:2 interactions with chain A- Metal complexes: A:H.41, A:C.145
QMEANDisCo Local
QMEAN Z-Scores
Template
2z9j.1.A
3C-like proteinase
Complex structure of SARS-CoV 3C-like protease with EPDTC
Complex structure of SARS-CoV 3C-like protease with EPDTC
- Seq Identity
- 96.08%
- Coverage
| Biounit Oligo State | Homo-dimer |
| QSQE | 0.95 |
| Method | X-ray, 1.95 Å |
| Seq Similarity | 0.62 |
| Coverage | 1.00 |
| Range | 1-302 |
Model-Template Alignment
Model 01
- Ligands
1 x GLY ,2 x O6K 1 x GLYCINEGLY.1: 3 residues within 4Å:- Chain A: R.298, S.301,
- Chain B: S.123
1 PLIP interactions:1 interactions with chain A- Hydrogen bonds: A:S.301
2 x ~{tert}-butyl ~{N}-[1-[(2~{S})-3-cyclopropyl-1-oxidanylidene-1-[[(2~{S},3~{R})-3-oxidanyl-4-oxidanylidene-1-[(3~{S})-2-oxidanylidenepyrrolidin-3-yl]-4-[(phenylmethyl)amino]butan-2-yl]amino]propan-2-yl]-2-oxidanylidene-pyridin-3-yl]carbamateO6K.2: 18 residues within 4Å:- Chain A: T.26, H.41, M.49, F.140, L.141, N.142, G.143, S.144, C.145, H.163, H.164, M.165, E.166, L.167, H.172, D.187, Q.189,
- Chain B: S.1
11 PLIP interactions:10 interactions with chain A, 1 interactions with chain B- Hydrophobic interactions: A:N.142, A:M.165, A:D.187, A:Q.189
- Hydrogen bonds: A:G.143, A:S.144, A:C.145, A:H.164, A:E.166, A:E.166, B:S.1
O6K.3: 18 residues within 4Å:- Chain A: S.1,
- Chain B: T.26, H.41, M.49, F.140, L.141, N.142, G.143, S.144, C.145, H.163, H.164, M.165, E.166, L.167, P.168, D.187, R.188
10 PLIP interactions:10 interactions with chain B- Hydrophobic interactions: B:N.142, B:M.165, B:E.166
- Hydrogen bonds: B:G.143, B:S.144, B:S.144, B:C.145, B:H.164, B:E.166, B:E.166
QMEANDisCo Local
QMEAN Z-Scores
Template
6y2g.1.A
3C-like proteinase
Crystal structure (orthorhombic form) of the complex resulting from the reaction between SARS-CoV-2 (2019-nCoV) main protease and tert-butyl (1-((S)-1-(((S)-4-(benzylamino)-3,4-dioxo-1-((S)-2-oxopyrrolidin-3-yl)butan-2-yl)amino)-3-cyclopropyl-1-oxopropan-2-yl)-2-oxo-1,2-dihydropyridin-3-yl)carbamate (alpha-ketoamide 13b)
Crystal structure (orthorhombic form) of the complex resulting from the reaction between SARS-CoV-2 (2019-nCoV) main protease and tert-butyl (1-((S)-1-(((S)-4-(benzylamino)-3,4-dioxo-1-((S)-2-oxopyrrolidin-3-yl)butan-2-yl)amino)-3-cyclopropyl-1-oxopropan-2-yl)-2-oxo-1,2-dihydropyridin-3-yl)carbamate (alpha-ketoamide 13b)
- Seq Identity
- 100.00%
- Coverage
| Biounit Oligo State | Homo-dimer |
| QSQE | 1.00 |
| Method | X-ray, 2.20 Å |
| Seq Similarity | 0.63 |
| Coverage | 1.00 |
| Range | 1-301 |
| Ligand | Added to Model | Description |
|---|---|---|
| ✓ | GLYCINE | |
| ✓ | ~{tert}-butyl ~{N}-[1-[(2~{S})-3-cyclopropyl-1-oxidanylidene-1-[[(2~{S},3~{R})-3-oxidanyl-4-oxidanylidene-1-[(3~{S})-2-oxidanylidenepyrrolidin-3-yl]-4-[(phenylmethyl)amino]butan-2-yl]amino]propan-2-yl]-2-oxidanylidene-pyridin-3-yl]carbamate | |
| ✓ | ~{tert}-butyl ~{N}-[1-[(2~{S})-3-cyclopropyl-1-oxidanylidene-1-[[(2~{S},3~{R})-3-oxidanyl-4-oxidanylidene-1-[(3~{S})-2-oxidanylidenepyrrolidin-3-yl]-4-[(phenylmethyl)amino]butan-2-yl]amino]propan-2-yl]-2-oxidanylidene-pyridin-3-yl]carbamate |
Model-Template Alignment
Model 03
- Ligands
2 x AZP 2 x (5S,8S,14R)-ETHYL 11-(3-AMINO-3-OXOPROPYL)-8-BENZYL-14-HYDROXY-5-ISOBUTYL-3,6,9,12-TETRAOXO-1-PHENYL-2-OXA-4,7,10,11-TETRAAZAPENTADECAN-15-OATEAZP.1: 23 residues within 4Å:- Chain A: T.25, T.26, H.41, M.49, Y.54, F.140, L.141, N.142, G.143, S.144, C.145, H.163, H.164, M.165, E.166, P.168, H.172, D.187, R.188, Q.189, T.190, A.191, Q.192
15 PLIP interactions:15 interactions with chain A- Hydrophobic interactions: A:H.41, A:M.49, A:P.168, A:Q.189
- Hydrogen bonds: A:F.140, A:N.142, A:G.143, A:S.144, A:S.144, A:C.145, A:H.164, A:E.166, A:E.166, A:Q.189
- Salt bridges: A:H.41
AZP.4: 23 residues within 4Å:- Chain B: T.25, T.26, H.41, M.49, Y.54, F.140, L.141, N.142, G.143, S.144, C.145, H.163, H.164, M.165, E.166, P.168, H.172, D.187, R.188, Q.189, T.190, A.191, Q.192
16 PLIP interactions:16 interactions with chain B- Hydrophobic interactions: B:H.41, B:M.49, B:P.168, B:Q.189
- Hydrogen bonds: B:F.140, B:N.142, B:G.143, B:S.144, B:S.144, B:C.145, B:H.164, B:E.166, B:E.166, B:E.166, B:Q.189
- Salt bridges: B:H.41
QMEANDisCo Local
QMEAN Z-Scores
Template
2a5i.1.A
3C-like peptidase
Crystal structures of SARS coronavirus main peptidase inhibited by an aza-peptide epoxide in the space group C2
Crystal structures of SARS coronavirus main peptidase inhibited by an aza-peptide epoxide in the space group C2
- Seq Identity
- 96.08%
- Coverage
| Biounit Oligo State | Homo-dimer |
| QSQE | 0.92 |
| Method | X-ray, 1.88 Å |
| Seq Similarity | 0.62 |
| Coverage | 1.00 |
| Range | 1-306 |
| Ligand | Added to Model | Description |
|---|---|---|
| ✓ | (5S,8S,14R)-ETHYL 11-(3-AMINO-3-OXOPROPYL)-8-BENZYL-14-HYDROXY-5-ISOBUTYL-3,6,9,12-TETRAOXO-1-PHENYL-2-OXA-4,7,10,11-TETRAAZAPENTADECAN-15-OATE | |
| ✓ | (5S,8S,14R)-ETHYL 11-(3-AMINO-3-OXOPROPYL)-8-BENZYL-14-HYDROXY-5-ISOBUTYL-3,6,9,12-TETRAOXO-1-PHENYL-2-OXA-4,7,10,11-TETRAAZAPENTADECAN-15-OATE | |
| ✕ - Not biologically relevant. | 1,2-ETHANEDIOL | |
| ✕ - Not biologically relevant. | 1,2-ETHANEDIOL | |
| ✕ - Not biologically relevant. | GLYCEROL | |
| ✕ - Not biologically relevant. | GLYCEROL |
Model-Template Alignment
Delete Model - ""
Are you sure you want to delete this model?
(This really can't be undone!)
Delete Project - ""
Are you sure you want to delete this project?
(This really can't be undone!)